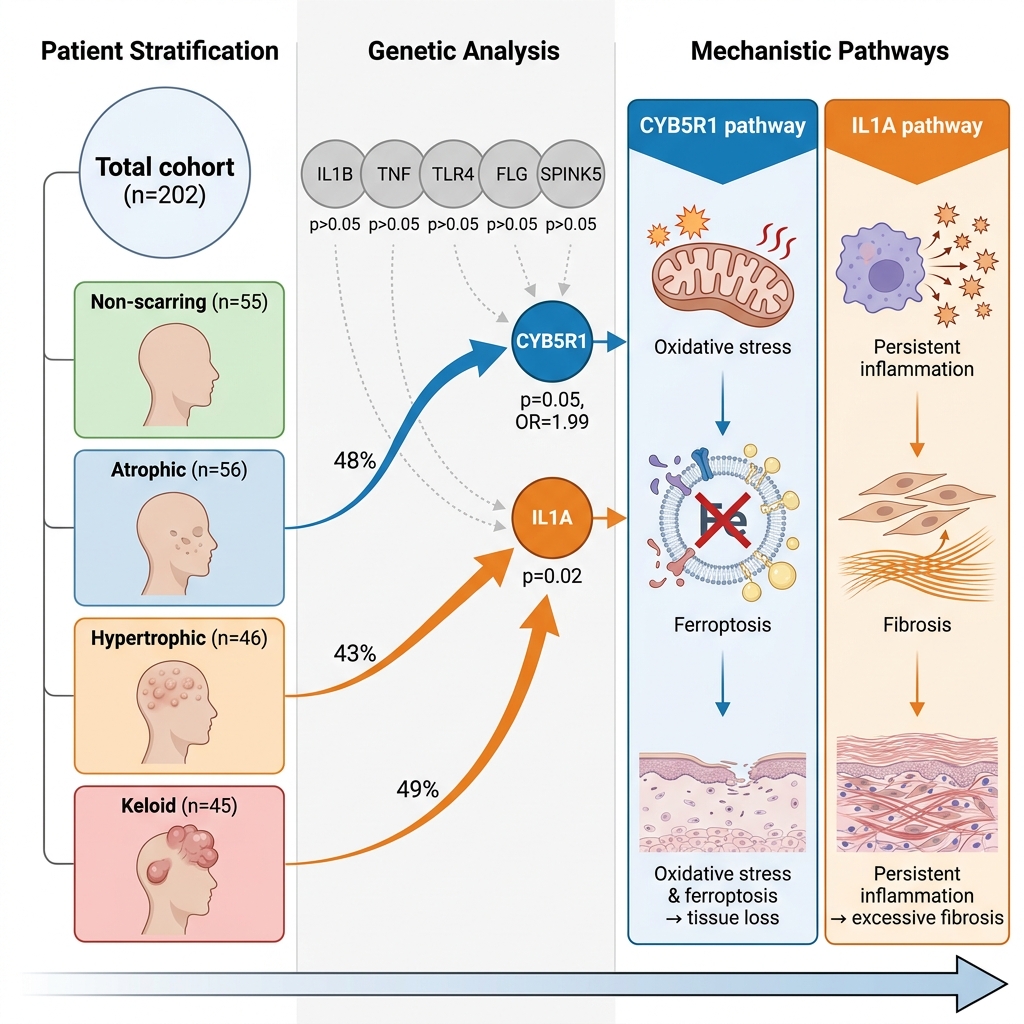
Potential genetic associations of acne scar phenotypes: IL1A in fibrotic scarring and CYB5R1 in atrophic scarring
- University of Health Sciences, Vietnam National University, Ho Chi Minh City, Viet Nam
- Vinmec Ocean Park 2 Hospital, Vinmec Healthcare System, Hung Yen, Vietnam
Abstract
Introduction: Nearly half of individuals diagnosed with acne vulgaris develop scarring. This study aimed to investigate the genetic associations between acne scarring and six selected gene variants (CYB5R1, IL1A, TLR4, FLG, SPINK5, and TNF).
Methods: We conducted a cross-sectional study at the University of Medicine and Pharmacy Hospital and the Vietnam National Hospital of Dermatology and Venereology from September 2023 to December 2024. A total of 202 acne patients were recruited and categorized into either a non-scarring cohort (n = 55) or a scarring cohort (n = 147). Patients with scars were further classified into atrophic (n = 56), hypertrophic (n = 46), and keloid (n = 45) subgroups.
Results: No significant demographic or clinical differences were observed between the groups (p > 0.05). Variants in CYB5R1 were more frequent in the scarred group than in the non-scarred group (38.1% vs. 23.6%, p = 0.05; OR = 1.99, 95% CI: 1.00–4.15), with the highest proportion observed in patients with atrophic scars. In contrast, IL1A variants were enriched in hypertrophic and keloid (fibrotic) scars (p < 0.05). The remaining evaluated genes (TNF, TLR4, FLG, and SPINK5) demonstrated no significant association with acne scarring.
Conclusion: This study suggests a potential association between CYB5R1 and atrophic scarring, as well as between IL1A and fibrotic scarring. These findings support the polygenic nature of acne scarring, although further validation in larger cohorts is required.
INTRODUCTION
Acne vulgaris is the eighth most common skin disease globally1 , affecting up to 85% of adolescents and often persisting into adulthood2. While inflammatory lesions may resolve, nearly half of affected individuals with acne suffer from develop scarring3 , which can lead to result in low self-esteem, social isolation, anxiety, and depression4. Scar formation can be broadly categorized divided into two primary phenotypes: those resulting from excessive tissue deposition, leading to hypertrophic or keloid scars, and those characterized by tissue loss or destruction, resulting in atrophic scars. Despite advances in acne treatment, the prevention and management of scarring remain therapeutic challenges hurdle3,5 , highlighting the critical need for research into acne scar morphogenesis and the early identification of individuals at elevated risk.
Current evidence indicates that acne scarring results from prolonged inflammation and dysregulated wound healing, where with genetic predisposition potentially plays may play a pivotal role3,5. Candidate genes implicated in inflammatory and immunological pathways, such as interleukin-1 (IL1), interleukin-6 (IL6), tumor necrosis factor (TNF), and toll-like receptor 4 (TLR4), have been associated with acne severity and inflammatory phenotypes6,7,8,9. Moreover, genes involved in skin barrier integrity and repair, including filaggrin (FLG) and serine protease inhibitor Kazal-type 5 (SPINK5), may influence scar formation by modulating epidermal homeostasis and inflammation resolution10,11. Additionally, the CYB5R1 gene, which encodes a member of the cytochrome b5 reductase family involved in oxidative stress and ferroptosis pathways, emerges as presents a novel candidate potentially affecting extracellular matrix remodeling in acne lesions12,13. Collectively, these six genes represent biologically plausible candidates for investigating the genetic basis of acne scarring.
However, despite growing interest in acne genetics, most research has targeted targets acne susceptibility rather than scarring outcomes6,7,8. To date, no comprehensive study has evaluated whether polymorphisms in IL1A, TLR4, TNF, FLG, SPINK5, and CYB5R1 predispose individuals to distinct acne scar phenotypes. Therefore, this study aims to investigate the correlation between these six genetic variants and acne scarring utilizing with detailed phenotypic scar characterization. By addressing this knowledge gap, our research seeks to enhance the understanding of acne scar pathogenesis and support the development of personalized approaches to in dermatological care.
MATERIALS AND METHODS
Patient Recruitment and Clinical Assessment
This cross-sectional study was conducted at the University of Medicine and Pharmacy Hospital, and the Vietnam National Hospital of Dermatology and Venereology between September, 2023 and December, 2024. The study protocol was approved by the Institutional Review Board of the School of Medicine (approval number #MEDVNU-093) and adhered to the principles of the Declaration of Helsinki.
A total of 202 patients with clinically diagnosed acne vulgaris were consecutively recruited during their outpatient dermatology visits. Eligible participants were at least 16 years of age, had a confirmed clinical diagnosis of acne vulgaris, and provided written informed consent. Patients with systemic inflammatory conditions, endocrine disorders, current immunosuppressive or systemic corticosteroid therapy, or a history of isotretinoin use within the preceding six months were excluded.
Participants were stratified into two groups based on their scar status: 55 patients with acne but without visible scars (the non-scarring cohort), and 147 patients with acne scars (the scarring cohort). Clinical characteristics, including age, sex, acne phenotype (comedonal, cystic, inflammatory, or mixed), and pore size (small, medium, or large), were recorded.
All patients were independently evaluated by two board-certified dermatologists, and any diagnostic discrepancies were resolved via consensus. Acne phenotypes were classified according to the criteria established by Sebastian Cruz et al. (2023)14. Scar assessment was performed by the same two board-certified dermatologists utilizing the morphological classification system (atrophic, hypertrophic, and keloid) as described by Jacob et al. (2001)15 and Kravvas et al. (2017)16. Additionally, scar severity was graded according to the Goodman & Baron qualitative global scarring grading system (2006)17 to ensure inter-rater reliability. Any further discrepancies were resolved by consensus. Clinical assessment and peripheral blood collection were performed concurrently to ensure phenotypic-genotypic concordance for downstream association analyses.
Sample Collection and DNA Extraction
Peripheral venous blood samples were collected in ethylenediaminetetraacetic acid (EDTA) tubes and processed according to standardized molecular laboratory protocols. Genomic DNA was extracted using validated commercial extraction kits. DNA concentration and purity were assessed via spectrophotometric and fluorometric methods. Samples that failed to meet quality control thresholds were excluded from subsequent analyses.
Single Nucleotide Polymorphism Selection and Targeted Sequencing
A targeted next-generation sequencing (NGS) approach was employed, focusing on preselected single nucleotide polymorphisms (SNPs) and pathogenic variants chosen based on biological plausibility and prior clinical evidence. The six selected gene variants (CYB5R1, IL1A, TLR4, FLG, SPINK5, and TNF) represent three principal biological axes implicated in acne pathogenesis: the inflammatory response, epidermal barrier function, and lipid metabolism.
Variant selection was informed by published evidence from the ClinVar and dbSNP databases, pathogenicity classification according to the American College of Medical Genetics and Genomics (ACMG) guidelines, and putative associations with inflammatory phenotypes, nodulocystic acne, and scar formation (Table 1). For variants without assigned reference SNP cluster IDs (rsIDs), genomic coordinates and Human Genome Variation Society (HGVS) nomenclature are provided to ensure reproducibility. High-depth NGS was performed to ensure the reliable detection of both rare and common variants within the targeted loci.
Selected gene variants information
| Gene variants | rsID | Database |
|---|---|---|
|
rsID: Not available IL1A(NM_000575.5):c.586C>T p.(Gln196Ter) |
ACMG: PVS1, PM2, PP3 | |
|
rs281865419 (NM_000594.4(TNF):c.322C>T (p.Arg108Trp)) | ClinVar, Pathogenic | |
|
rsID: Not available TLR4(NM_138557.3):c.1390del p.(Cys464AlafsTer2) |
ACMG: PVS1, PM2 | |
|
rs771730802 (NM_006846.4(SPINK5):c.81+5G>A)(intron variant) | ClinVar, Pathogenic | |
|
rs775253166 NM_002016.2(FLG):c.11033_11036del (p.Ser3678fs) | ClinVar, Pathogenic | |
|
rsID: Not available CYB5R1(NM_016243.3):c.706C>T p.(Arg236Cys) CYB5R1(NM_016243.3):c.624C>A p.(Cys208Ter) |
ACMG: PVS1, PM2 |
Genetic Analysis
Genetic analysis was designed based on recent genome-wide association studies (GWAS) of acne vulgaris, which have identified more than 25 susceptibility loci across diverse populations8,18. To capture the genetic architecture relevant to acne and scarring, we selected two complementary SNP panels. First, we compiled index SNPs and their proxies (r² ≥ 0.8) reported in large meta-GWAS of acne vulgaris (including loci near FADS2, LGR5, FASN, ZNRF3-KREMEN1, HLA-C, and FLG)8,18. These SNPs represent established risk loci implicated in sebaceous gland biology, hair follicle development, immune regulation, and wound healing.
Second, to expand genomic coverage, we included a custom array panel of approximately 4,500 SNPs, incorporating variants previously reported in dermatologic genetics studies and polymorphisms in inflammation-related, barrier-related, and extracellular matrix remodeling genes6,19. This panel was designed to facilitate exploratory association testing beyond the known GWAS hits, with a specific focus on genes potentially involved in scarring.
Genotyping was performed using the Illumina Infinium Global Screening Array (GSA) supplemented with custom content, with quality control assessments conducted by Macrogen Inc. (Seoul, Korea). In brief, DNA samples (≥50 ng/µL, A260/280 ratio 1.8–2.0) were processed according to the manufacturer's protocols. Quality control procedures followed standard GWAS pipelines20. Specifically, samples with call rates <95%, sex discordance, or cryptic relatedness (pi-hat >0.2) were excluded. SNP-level filters included a call rate <98%, a Hardy–Weinberg equilibrium of p < 1 × 10⁻⁶, and a minor allele frequency (MAF) <0.01. After quality control, approximately 4,300 SNPs remained for downstream analysis.
Statistical association analyses were conducted using PLINK v1.9 software. Additive genetic models were tested using logistic regression, adjusted for age, sex, and acne severity. Principal component analysis (PCA) was employed to control for population stratification, and the first five principal components (PCs) were included as covariates. Variants showing a suggestive association (p < 1 × 10⁻⁴) were further annotated using ANNOVAR against the dbSNP, gnomAD, and GTEx eQTL databases. Enrichment analyses of significant SNPs were performed to identify biological pathways relevant to scar pathogenesis.
Statistical Analyses
IBM SPSS Statistics version 26.0 and R version 4.5.1 were utilized for the statistical analyses. Independent t-tests were used to compare continuous variables between groups, and chi-square tests were used to assess categorical relationships, with statistical significance set at p < 0.05. The R programming language was used to visualize gene variant distributions and odds ratios using forest plots and heatmaps.
RESULTS
Demographic Characteristics and Prevalence of Acne Scarring
Over a 16-month period, the study recruited 202 patients receiving acne treatment at the Vietnam National Hospital of Dermatology and Venereology. The participants were stratified into two groups based on scar status: a non-scarring cohort (n = 55) and a scarring cohort (n = 147). Subsequently, gene analysis was conducted, and clinical features were assessed by dermatologists. Detailed data are presented in Table 2.
Demographic characteristics of the study groups
| No scar (n=55) | Scar (n=147) | p-value | |
|---|---|---|---|
| Gender | |||
| Female | 23 (41.8) | 72 (49.0) | 0.454 |
| Male | 32 (58.2) | 75 (51.0) | |
| Age, years | |||
| Mean (SD) | 29.3 (7.00) | 28.4 (7.33) | 0.435 |
| Median (IQR) | 28 (17 - 42) | 28 (17 - 42) | |
| Acne phenotypes (%) | |||
| Comedonal | 18 (32.7) | 40 (27.2) | 0.883 |
| Cystic | 12 (21.8) | 33 (22.4) | |
| Inflammatory | 12 (21.8) | 34 (23.1) | |
| Mixed | 13 (23.6) | 40 (27.2) | |
| Pore size (%) | |||
| Small | 26 (47.3) | 55 (37.4) | 0.246 |
| Medium | 18 (32.7) | 46 (31.3) | |
| Large | 11 (20.0) | 46 (31.3) | |
| Found | 19 (34.5) | 37 (25.2) | 0.185 |
| None | 36 (65.5) | 110 (74.8) | |
| Found | 14 (25.5) | 46 (31.3) | 0.419 |
| None | 41 (74.5) | 101 (68.7) | |
| Found | 18 (32.7) | 54 (36.7) | 0.597 |
| None | 37 (67.3) | 93 (63.3) | |
| Found | 13 (23.6) | 56 (38.1) | 0.05 |
| None | 42 (76.4) | 91 (61.9) | |
| Found | 13 (23.6) | 50 (34.0) | 0.156 |
| None | 42 (76.4) | 97 (66.0) | |
| Found | 24 (43.6) | 46 (31.3) | 0.101 |
| None | 31 (56.4) | 101 (68.7) | |
No significant differences were observed between the groups regarding gender distribution, age, and clinical features (p > 0.05). Genetic analysis revealed no significant differences in the proportion of variants in SPINK5, TNF, IL1A, FLG, and TLR4 (p > 0.05) between the non-scarring and scarring cohorts. In contrast, CYB5R1 variants, though not statistically significant, demonstrated a borderline association with the scarring cohort (p = 0.05).
Genetic Associations with Scarring and Non-Scarring Outcomes
This genetic association study investigated the relationship between scarring risk and selected gene variants (Figure 1). Notably, CYB5R1 variants exhibited the strongest observed association with scarring (OR = 1.99, 95% CI: 1.00 - 4.15); however, because the confidence interval includes unity, this finding does not reach conventional statistical significance.
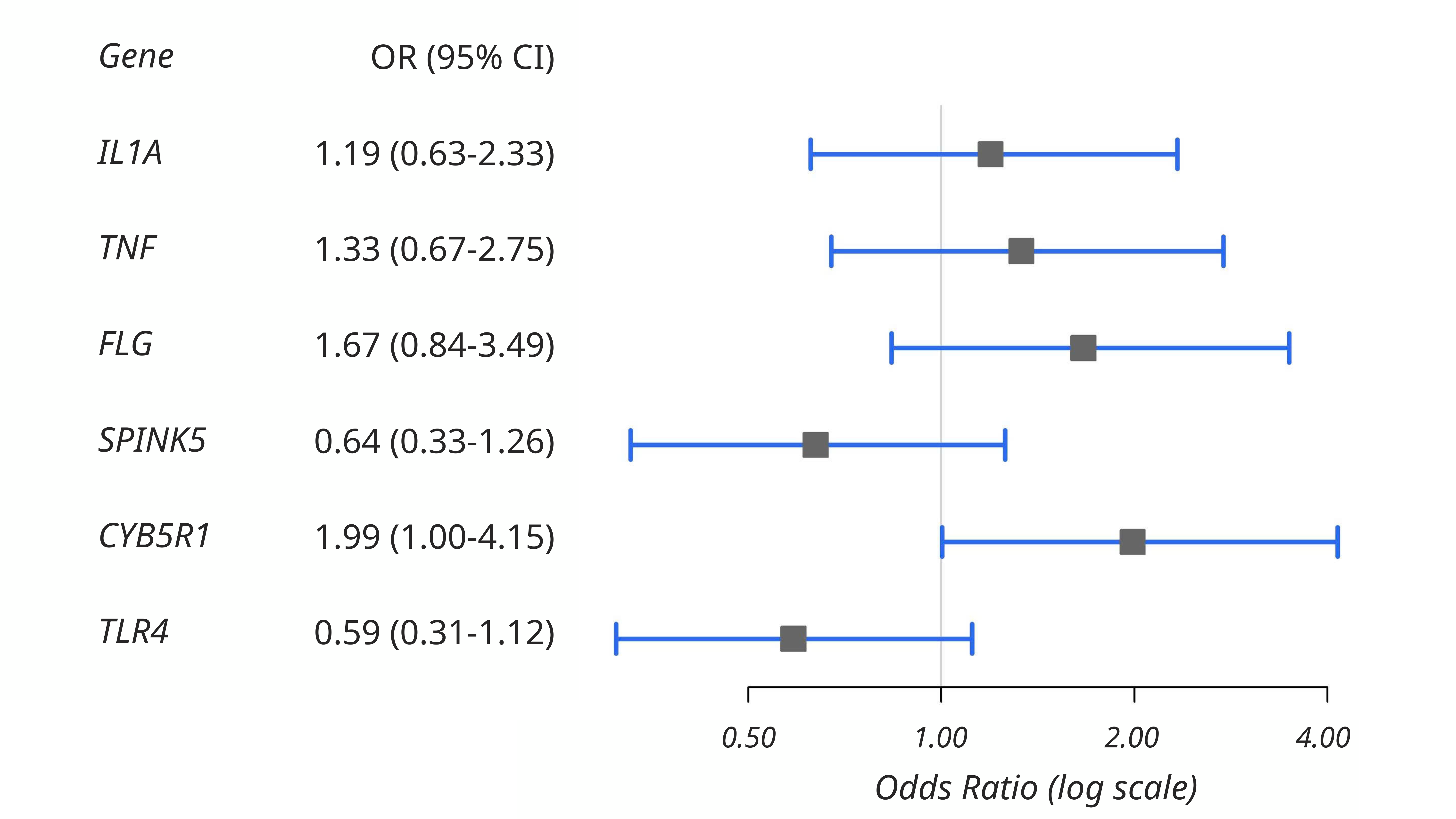
Association of genetic variants with the risk of acne scarring. This forest plot illustrates the odds ratios (OR) and 95% confidence intervals (CI) for the association between acne scarring and variants in the
Although FLG, IL1A, and TNF variants showed odds ratios greater than 1, their wide confidence intervals overlapping with unity indicated a lack of statistical significance. Conversely, TLR4 (OR = 0.59, 95% CI: 0.31 - 1.12) and SPINK5 (OR = 0.64, 95% CI: 0.33 - 1.26) were associated with reduced odds, but these findings were also not statistically significant.
Genetic Associations with Specific Scar Phenotypes
Participants in the scarring cohort were stratified into three scar types of increasing severity: atrophic (n = 56), hypertrophic (n = 46), and keloid (n = 45). The study examined the odds ratios and the distribution of gene variants across these three scar types. Detailed data are presented in Table 3. Significant differences in variant proportions were observed across scar types for IL1A (p = 0.020) and CYB5R1 (p = 0.042). No significant differences were found for TLR4, FLG, SPINK5, or TNF (p > 0.05) across the scar phenotypes.
Distribution of Gene Variants Across Diverse Scar Phenotypes
| Gene Variants | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Found (n=72) | None | Found (n=70) | None | Found (n=63) | None | Found (n=56) | None | Found (n=69) | None | Found (n=60) | None | |
|
No Scars (n = 55) | 18 | 37 | 24 | 31 | 13 | 42 | 19 | 36 | 13 | 42 | 14 | 41 |
|
Atrophic (n = 56) | 12 | 44 | 20 | 36 | 16 | 40 | 16 | 40 | 27 | 29 | 18 | 38 |
|
Hypertrophic (n = 46) | 20 | 26 | 11 | 35 | 18 | 28 | 11 | 35 | 16 | 30 | 17 | 29 |
|
Keloid (n = 45) | 22 | 23 | 15 | 30 | 16 | 29 | 10 | 35 | 13 | 32 | 11 | 34 |
| p-value | 0.020* | 0.225 | 0.335 | 0.511 | 0.042* | 0.495 | ||||||
The proportions of patients harboring specific gene variations across various scar types were illustrated via heatmap analysis (Figure 2). The proportion of the CYB5R1 gene variant in atrophic scars was found to be 0.48. The proportion of the IL1A gene variant was 0.43 in hypertrophic scars and 0.49 in keloid scars. In patients without scars, the proportion of the TLR4 gene variant was 0.44. Hypertrophic scars demonstrated an FLG gene variant proportion of 0.39, while keloid scars showed a variant proportion of 0.36. In patients without scars, the proportion of the SPINK5 gene variant was 0.35. With values of 0.32 in atrophic scars, 0.37 in hypertrophic scars, 0.24 in keloid scars, and 0.25 in the non-scarring cohort, the TNF gene variant exhibited moderate proportions across all groups.
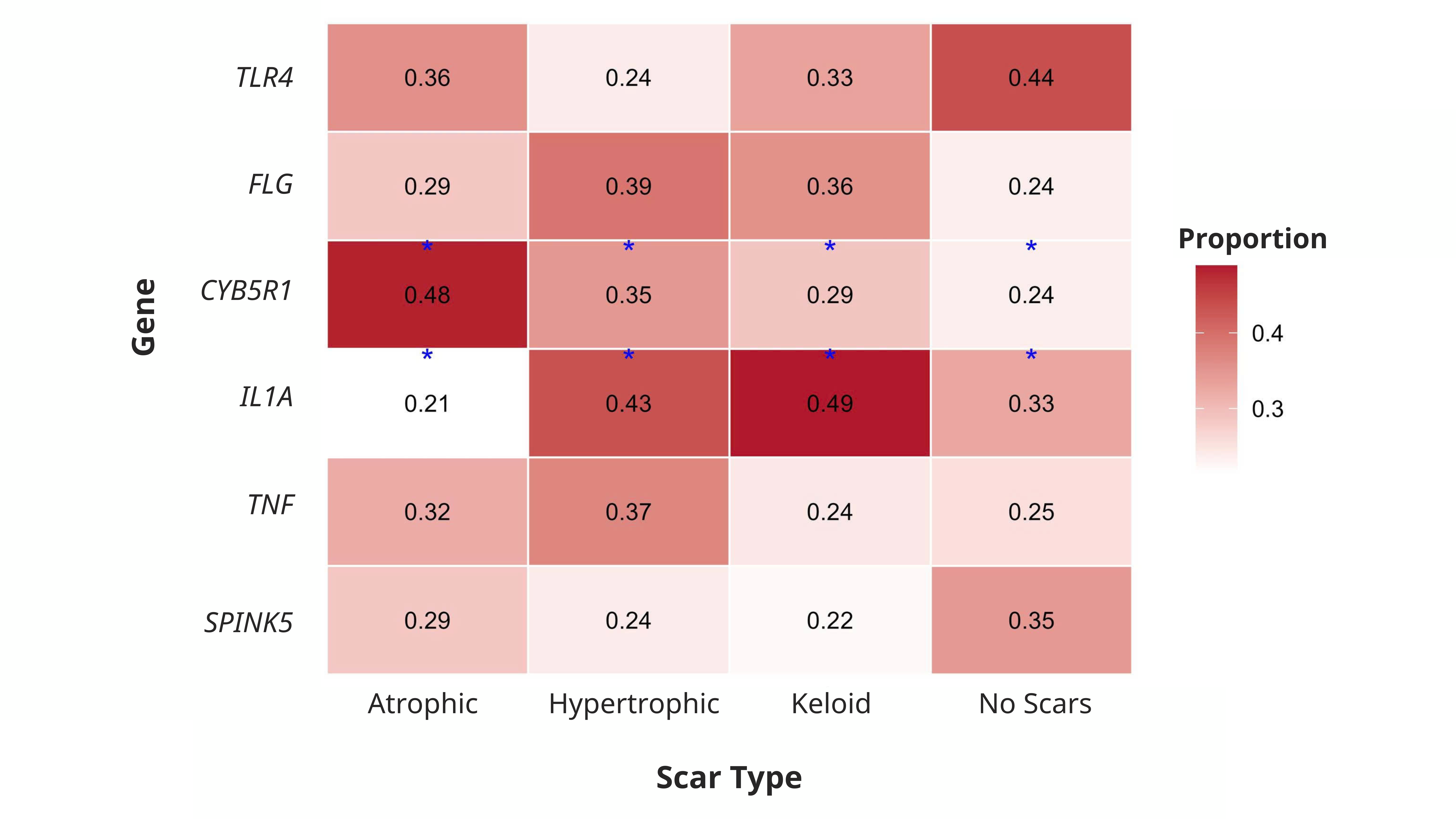
Proportion of gene variants acrossacnescar phenotypes. This heatmap illustrates the proportion of individuals in the atrophic, hypertrophic, keloid, and non-scarring cohorts who harbored variants in the
DISCUSSION
This study found no significant associations between demographic factors (gender, age), clinical acne phenotypes, or pore size and the presence of acne scarring. These findings align with previous reports indicating that such variables often have limited predictive value for scarring outcomes. Moreover, we observed a potential association between CYB5R1 variants and acne scarring, highlighting a possible genetic predisposition to this condition. This variant is more prevalent in the scarred cohort, presenting mostly with atrophic scars, and carries nearly double the odds of scarring (OR 1.99; CI 1.00 - 4.15), while the other gene variants showed no significant association. However, this finding was borderline and should be interpreted cautiously.
According to meta-analyses, between 75 and 90 percent of acne scars are atrophic3,21, with hypertrophic and keloidal scars accounting for only a small minority. Indeed, Cheng et al. discovered that approximately 78% of scars in acne patients were atrophic3, consistent with older estimates of 80-90%21. Therefore, genes that affect the development of atrophic scars are of significant clinical relevance in acne pathogenesis.
CYB5R1 encodes NADH-cytochrome b5 reductase 1, a ubiquitous oxidoreductase involved in lipid metabolism, steroid biosynthesis, and drug metabolism, with strong expression in keratinocytes and mitochondria across multiple tissues. Crucially, recent mechanistic work implicates CYB5R1 in the generation of reactive oxygen species (ROS) and lipid peroxidation—especially in the context of ferroptosis (iron-dependent lipid peroxidation)22. CYB5R1 is predicted to localize to the mitochondria and the endoplasmic reticulum and shares 63% sequence identity with the well-known CYB5R3 isoform. Recent data indicate that CYB5R1 can drive mitochondrial electron leakage to generate ROS12.
We hypothesize about the mechanisms underlying CYB5R1's impact on scarring given its redox involvement. Our findings are consistent with new research that links CYB5R1 to hydrogen peroxide production and membrane damage during ferroptotic cascades22, pointing to a mechanism by which its variants increase the oxidative burden in acne-prone skin. Although ferroptosis is known to trigger inflammatory pathways, an overproduction of ROS may affect the viability or function of dermal fibroblasts, resulting in inadequate collagen deposition and cutaneous atrophy22. Likewise, fibroblasts or endothelial cells in the regenerating skin may directly undergo cell death as a result of CYB5R1-mediated ferroptosis. On the other hand, hypertrophic/keloid scars are indicative of increased matrix and fibroblast activity; hence, a pro-oxidative bias may specifically predispose individuals to the atrophic phenotype. The association of CYB5R1 adds a new redox-mediated dimension to scar morphogenesis, which is distinct from well-established genes like IL1A and TNF, which are well-documented in acne pathogenesis23. Future research should examine whether individuals harboring the CYB5R1 variant actually exhibit elevated oxidative stress or compromised fibroblast remodeling during the healing of acne lesions. Nevertheless, despite its role in regulating ferroptosis and oxidative stress—processes essential to lipid peroxidation and cellular death12,23—CYB5R1 has not previously been studied in dermatological settings like acne or skin fibrosis.
The cytokines of the IL-1 family are well-known mediators of acne inflammation and scarring. Historically, IL1A had been found to be particularly important because IL-1α activity is detected early in comedogenesis and can help initiate the inflammatory response in acne lesions24. Previously, IL-1 protein levels were shown to be significantly higher in fibrotic scars, such as keloids, which indicates persistent wound inflammation25; however, previous genetic research did not find any link to acne scarring26. Consequently, our discovery of a significant IL1A gene variant association (p=0.020) offers compelling evidence that IL-1α signaling affects scar risk. Interestingly, IL1B was previously recognized as a major cytokine in acne26; subsequently, our findings are consistent with known biology, but they also point to a new polymorphism or regulatory mechanism in the IL1A gene that was missed by previous SNP investigations.
TNF-α is also a key inflammatory mediator in acne. In keloid tissues, the TNF-α protein is increased, and TNF-α mediates the immediate acne lesion response25. However, SNPs in the TNF promoter, such as -308G/A, have not reliably predicted the severity or amount of scarring26,27. All scar types in our group had modest frequencies (0.24–0.37) of TNF variants. This is consistent with the complex body of evidence, as meta-analyses only find minor, population-specific effects of TNF SNPs on acne. Though its genetic contribution to scarring seems widespread and most likely involves numerous regulatory alleles rather than a single common variant, TNF's key inflammatory role remains evident overall25,26.
For TLR4, the pattern was intriguing. TLR4 is an innate immune receptor that initiates MyD88/NF-κB signaling, which causes dermal fibroblasts to produce IL-6, IL-8, and MMP-128. In fact, fibroblasts exposed to UV-A exhibit elevated TLR4 expression and subsequent production of IL-6, IL-8, and MMP-128. The importance of 3′-UTR SNPs in TLR4 linked to acne susceptibility has been confirmed by recent GWAS on acne7. According to our study, the no-scar group had the highest proportion of the TLR4 variant (44%), indicating a protective polymorphism that inhibits fibrotic/inflammatory signaling. This observation aligns with biological mechanisms; for instance, in skin models, inhibiting TLR4 reduces pro-fibrotic cytokines. Therefore, by suggesting that specific alleles may reduce scar development, our findings expand on TLR4's recognized role in acne7,28.
Furthermore, there were notable patterns in the genes related to skin barrier function. Filaggrin, or FLG, is necessary for epidermal cornification and hydration; ichthyosis and severe eczema result from null variants29. We found FLG variants in approximately 36–39% of hypertrophic and keloid scars, despite the fact that they were previously considered to be unrelated to overall acne risk. According to this new association, barrier abnormalities may make people more likely to experience excessive fibrosis during the healing process. Similarly, SPINK5 produces the protease inhibitor LEKTI, which is necessary for cornification; variants in SPINK5 result in Netherton syndrome, which is characterized by scaly, irritated skin30,31. Interestingly, the SPINK5 variant was enriched in those without scars (35%), suggesting that some SPINK5 alleles may reduce inflammation that promotes scarring. These findings raise novel questions about how barrier gene variation can affect scar outcomes, because FLG and SPINK5 polymorphisms have not, to our knowledge, been connected to acne scarring previously29.
This study has several strengths, including its detailed phenotypic characterization of acne scars (atrophic, hypertrophic, and keloid subtypes) by board-certified dermatologists using standardized classification systems, which enhances the clinical relevance of the findings. The use of a custom-expanded genotyping array allowed for the exploration of biologically plausible candidate genes beyond established GWAS loci, providing insights into potential redox-mediated mechanisms in acne scarring pathogenesis. Additionally, the cross-sectional design enabled efficient recruitment and simultaneous assessment of genetic variants and scar outcomes in a Vietnamese hospital-based cohort, where such data remain limited. However, the relatively small sample size, particularly within the scar subgroups (n = 202 overall, with scar subgroups of n = 45-56), reduces statistical power and increases the risk of type II errors. The primary association for CYB5R1 variants was borderline (p = 0.050, OR = 1.99, 95% CI: 1.00-4.15), rendering it nominally significant and exploratory rather than confirmatory. The cross-sectional design prevents causal inference, and findings require replication in larger, independent cohorts for confirmation and generalizability.
Conclusion
In conclusion, our findings suggest a possible distinct role for CYB5R1 and IL1A variants in atrophic and fibrotic scarring, respectively. Hypothetical mechanisms are suggested to involve oxidative stress and ferroptosis pathways, which represents a paradigm-shifting novelty, connecting redox biology with acne pathophysiology, even though the findings regarding TNF, FLG, SPINK5, and TLR4 collectively build upon established inflammatory and barrier pathways in dermatology. These discoveries support the polygenic character of acne scarring and open the door for tailored treatments, such as ferroptosis inhibitors, to lessen scar formation in people who are genetically predisposed.
ABBREVIATIONS
CYB5R1: Cytochrome B5 reductase 1; SPINK5: Serine peptidase inhibitor Kazal type 5; FLG: Filaggrin; TNF: Tumor necrosis factor; IL: Interleukin; SNP: Single Nucleotide Polymorphisms; ROS: Reactive Oxygen Species; NGS: Next-generation sequencing; GWAS: Genome-wide association studies; PCA: Principal component analysis; OR: Odds ratio; CI: Confidence Interval; SD: Standard deviation; IQR: Interquartile range
ACKNOWLEDGEMENTS
We would like to convey our heartfelt thanks to the physicians and clinical personnel of the University of Medicine and Pharmacy Hospital and the Vietnam National Hospital of Dermatology and Venereology for their outstanding assistance and collaboration during this study.
AUTHOR CONTRIBUTIONS
Conceptualisation: Duy-Khang Nguyen, Quynh-Giang Nguyen, Dong Kha To. Data analysis and methodology: Duy-Khang Nguyen, Quynh-Giang Nguyen, Chi-Bao Bui. Acquisition of clinical data: Tuan-Khoi Duong, Dang-Khoa Nguyen, Dong Kha To. Writing - original draft preparation: Duy-Khang Nguyen, Quynh-Giang Nguyen, Hoang-Dung Nguyen. Writing - review & editing: Duy-Khang Nguyen, Chi-Bao Bui, Quynh-Giang Nguyen, Dong Kha To. Supervision: Chi-Bao Bui, Dong Kha To. All authors have read and agreed to the published version of the manuscript.
COMPETING INTEREST
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
ETHICS APPROVAL
The study protocol was approved by the Institutional Review Board of School of Medicine (approval number #MEDVNU-093) and adhered to the principles of the Declaration of Helsinki.
FUNDING
This study was primarily funded by the Vietnam National University Ho Chi Minh City (#C2024-44-24).
DATA AVAILABILITY STATEMENT
On reasonable request, the associated author will provide the data used in this article.
CONSENT FOR PUBLICATION
Not applicable.

